Follow us on Twitter

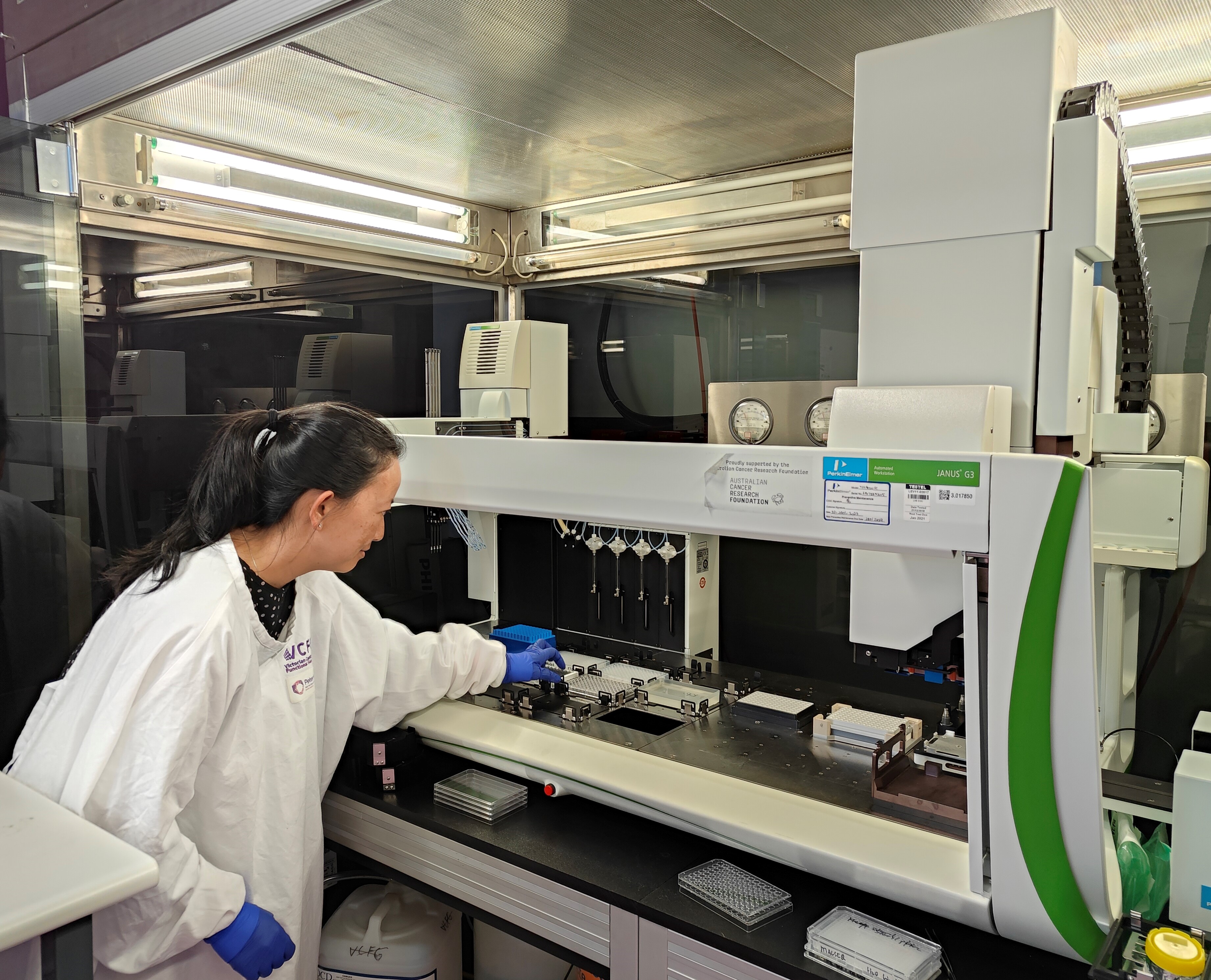
The Victorian Centre for Functional Genomics (VCFG) at Peter Mac is a national platform that enables biomedical researchers to perform novel discovery-based high throughput screens using multiple platforms. The Victorian Centre for Functional Genomics facilitates compound, CRISPR and RNAi screening at any scale in both 2D and 3D formats, using liquid handling instruments, automated high content imaging microscopes, live imaging systems and large reagent libraries.
The Victorian Centre for Functional Genomics provides a collaborative and innovative partnership. Our most common operations model is ‘researcher driven, staff assisted’ model where you embed in our lab and be trained on the appropriate instruments to perform your screen under the guidance of our team. We also provide ‘fee for service’ projects when we have team capacity, at the whole project level or a hybrid version that works for you. All data generated remains the intellectual property of the researcher. Importantly, each project is customised to your specific biological question, and we will help drive your project to the best screen outcome possible. The partnership beings with a discussion with Associate Professor Kaylene Simpson and a team of highly experienced research team.
Victorian Centre for Functional Genomics platforms and resources
We have a diverse portfolio of research platforms, many of which can have substantial overlap, depending on the project. Each platform has an overarching user guide and a suite of SOPs that are frequently updated. You will be given the appropriate documents when you first connect with us to discuss a project, and before you get trained on any instrumentation.
Our data analysis and machine learning platform is the cornerstone of the outputs generated from all the screens performed with us. Our key strength is in quantitative cellular phenotyping of cells grown in 2D and spheroids and organoids following perturbation in arrayed screening formats. We have developed sophisticated and highly customisable analysis pipelines by a team of specialised analysts, enabling us to assist you in interpreting the response of morphological perturbation in the cells using an unbiased method, dissect cellular responses and make therapeutic predictions (Choo et al 2021; Ramm et al 2022).
For a more details, see our short explainer.
The Victorian Centre for Functional Genomics has established a high throughput pipeline for embedding cells or organoids in Matrigel using our robotic automation or customisable hydrogel using our Rastrum bioprinter instrument from Inventia Life Science. This platform enables highly complex co-culture interaction studies, including tumour and immune cells, host-pathogen interactions, angiogenesis and cell invasion. For a broader overview and some FAQs see our short explainer.
Resources available:
- Matrigel (sourced in large volumes to provide consistent lot numbers)
- Hydrogel (Rastrum bioink)
With the Molecular Genomics Core we have recently developed a high throughput sequencing method called MAC-seq (Multiplexed Analysis of Cells). This can be performed in 384 well format in any experimental setting including time course analysis of growth or death, response to drug treatment and characterisation of patient materials in 3D matrices. Importantly, when coupled with high content imaging it allows us to perform 'Imaging Transcriptomics' further delineating our hit lists based on phenotypic quantitation together with target activity, see Li et al 2023. For a broader overview and some FAQs see our short explainer.
Using high throughput liquid handling automation, the Victorian Centre for Functional Genomics enables all types of compound screening approaches from cell-based to cell-free systems. We can deliver compounds in both 2D and 3D format and can accommodate dose curves with BYO drugs in multiple cell lines, 2- and 3-way drug synergy and large-scale collections. There is great creativity in this screening approach, and we work with you to develop the most appropriate experimental design and assay readouts.
Resources available:
- Accessed from Compounds Australia Griffith University (commonly screened libraries include the FDA, epigenetics, kinase inhibitor and scaffold collections, either as the whole collection or custom cherry picked)
- BYO compounds welcome
CRISPR screens enable researchers to determine resistance in their cellular model, facilitate biomarker discovery in patient samples or determine genes responsible for synthetic lethality. The Victorian Centre for Functional Genomics supports both synthetic and pooled CRISPR screening strategies. More details on this platform can be found below and in our short FAQ explainer.
Synthetic CRISPR
The synthetic CRISPR platform enables gene knockout in an arrayed format using liquid handling automation in 96- or 384-well format. This platform uses synthetic purified sgRNAs that are transiently delivered into Cas9-stable cells using lipid transfection reagents or nucleofected with purified Cas9 protein. The phenotype is assessed typically within 72-96 hours. This platform enables cell phenotyping approaches using high content imaging, fluorescence/luminescence plate reader end points or FACS profiling.
Resources available:
- Human whole genome arrayed sgRNA library (Horizon Discovery)
- Screen as a whole genome (61 plates) or cherry pick custom sub-sets
- This library is part of a 3-way venture between the Victorian Centre for Functional Genomics, ANU Centre for Therapeutic Discovery and Functional Genomics South Australia and supported by Phenomics Australia
- Arrayed synthetic single crRNA kinome
- Cas9 virus or protein
- Nucleofection reagents
- Purchasing sgRNAs at discounted rates
Viral CRISPR
The Victorian Centre for Functional Genomics houses both pooled and arrayed viral CRISPR libraries. Pooled CRISPR library are analysed using next generation sequencing. We take you through the screening process step-by-step for both whole genome pooled library or custom pooled sublibrary screens, from planning to execution through to analysis. The TransEDIT dual guide viral library in arrayed format can be used for single target validation, or boutique pooled library screening.
Resources available:
- CRISPR KO Brunello (human) and Brie (mouse) pooled libraries
- CRISPRa Calabrese (human) and Caprano (mouse) pooled libraries
- CRISPRi Docetto (human) and Dolomiti (mouse) pooled libraries
- Custom pooled CRISPR sublibrary generation and reagents
- Library amplification protocols and reagents
- In-house NGS (in collaboration with Molecular Genomics Core) and analysis pipelines
- Cas9 and dCas9 virus
- TransEDIT dual guide whole genome human arrayed viral library
The end point assay of an arrayed screen can simply look at changes in cell viability following perturbation or can be far more complex with phenotypic outputs detecting changes at sub-cellular levels. Cell painting can be undertaken utilising any combination of validated cellular machinery stains and we have optimised staining protocols for both 2D adherent cells and 3D spheroid and organoid cultures. These phenotypic changes are then measured using our wide range of instrumentation that allows both live and fixed cell imaging in bright field and multi-channel fluorescence using either widefield or confocal microscopy. Collectively, we have a high throughput instruments that can maximise the amount of information we can obtain from any screen.
Resources available:
The siRNA platform is based on Horizon Discovery's siGENOME SMARTpool reagents arrayed in either 96- or 384-well plate format. siRNAs targeting the protein coding genome, miRNAs (mimic or inhibitor) or long non-coding RNAs are transiently introduced into cells using lipid-based reagents and assayed between 72-144 hours post-transfection. This platform offers similar readout options as screening arrayed CRISPR.
The Victorian Centre for Functional Genomics has a pooled shRNA screening platform to enable stable knockdown in primary, non-dividing and standard cell lines. shRNAs can be screened singularly for validation purposes or screened as genome-wide or custom pools. The Victorian Centre for Functional Genomics together with the Molecular Genomics Core has developed the next-generation sequencing pipeline to identify the genes responsible for the biological phenotype of an assay.
Victorian Centre for Functional Genomics supporting services
VCFG PRoject Information and Management Enterprise (PRIME)
Every project is securely managed through PRIME, our project management portal using the Atlassian tool Confluence. Analysis is provided in a fully interactive HTML format with easy access to raw data, code and representative images. This portal future-proofs our capacity to revisit your dataset for publication, for re-analysis or for large-scale sharing. We also use this portal to collate funding and outputs (abstracts, presentations, publications) for future reporting.
Grants
We proudly support your grant applications through developing a customised approach to your specific project and budget. Happy to discuss your ideas! Email: This email address is being protected from spambots. You need JavaScript enabled to view it..
Partner organisations
The Victorian Centre for Functional Genomics is a partner of MACH OMICS. Visit the OMICS directory and collective mission.
Victorian Centre for Functional Genomics instrumentation
We have a large array of instrumentation available for anyone to access. We will provide you with instrument user guides prior to training.
- BioTek EL406: three units that aspirate and dispense media, cells, drugs and fix and stain
- BioTek Multi-flo: like the EL406 but can aspirate and dispense any plate type and has a little more nozzle position control
- Caliper ALH3000 liquid handling robot for all transfection and speciality liquid handling applications
- Perkin Elmer Janus G3 liquid handling robot
- Tecan D300E drug dispenser (excellent for BYO compounds, ultra-low dead volume, can randomise plate positions in a flash, super quick delivery)
- BioTek Cytation 5 plus BioSpa, a sophisticated plate imager and reader with live cell imaging
- BioTek Cytation C10, a sophisticated imager with spinning disk confocal and plate reader
- Fully automated Cellomics CX-7 LZR and CX-7 LZR Pro (2 units), laser light, wide field, confocal, live cell imaging
- Incucyte SX5 live cell imaging, metabolism module, wound healing x 2 instruments
- AMAXA 4D nucleofector and 96-well adaptor for high throughput
- Neon NxT electroporation system
- Barcode printer; all instruments are barcode-read enabled
- EVOS Fl, bench top fluorescent microscope
- IsoLight from Isoplexis
- LiCONICs 220 stacking tower incubator
- Rastrum 3D bioprinter
- Seahorse metabolic bioanalyser
Team members
Acknowledgements
The Victorian Centre for Functional Genomics has infrastructure funding and support from




Researchers accessing the Victorian Centre for Functional Genomics have funding from









Since 2008, the Victorian Centre for Functional Genomics has been a member of 4 successful ACRF infrastructure program applications


















